How Gut Bacterium Bt Unlocks Fiber's Hidden Nutrition

TL;DR: Scientists identified a specific sugar molecule (glucorhamnan) from the gut bacterium Ruminococcus gnavus that triggers Crohn's disease flares by activating TLR4 immune receptors. This breakthrough shifts IBD research from vague microbiome imbalance to a precise molecular target, opening paths for targeted diagnostics and treatments.

In 2019, a team led by Matthew Henke in Jon Clardy's laboratory at Harvard published findings in the Proceedings of the National Academy of Sciences that pinpointed an inflammatory polysaccharide produced by Ruminococcus gnavus, a bacterium that lives in virtually everyone's gut. This wasn't just another microbiome correlation study. The researchers isolated a specific molecule, a glucorhamnan-type polysaccharide made of glucose and rhamnose sugars, and showed it could directly activate immune cells called dendritic cells to pump out TNF-alpha, a potent inflammatory cytokine.
The mechanism was precise. The glucorhamnan binds to Toll-like receptor 4 (TLR4) on dendritic cells, triggering the same alarm system that normally responds to dangerous bacterial infections. As Henke put it, this is "a distinct molecule that represents the potential link between gut microbes and an inflammatory disease." The team even identified the gene cluster responsible for producing the glucorhamnan, opening a clear path toward targeted intervention.
"This is a distinct molecule that represents the potential link between gut microbes and an inflammatory disease."
- Matthew Henke, Harvard University / Clardy Laboratory
To appreciate why this matters, you need to understand how far microbiome science has come, and how stuck it was until recently.
The story of gut bacteria and disease goes back over a century. In the early 1900s, Elie Metchnikoff proposed that gut microbes influenced health and aging, though his ideas were largely dismissed by mainstream medicine. For most of the twentieth century, the gut microbiome was a black box. Doctors knew that Crohn's disease involved inflammation of the digestive tract, but the triggers remained mysterious.
The genomics revolution of the 2000s changed everything, sort of. Suddenly, researchers could sequence all the DNA in a stool sample and catalog thousands of bacterial species. Studies consistently found that Crohn's patients had altered microbiomes compared to healthy controls, a state called dysbiosis. R. gnavus kept showing up as particularly enriched during disease flares, sometimes becoming the most abundant species in the entire gut.

But correlation isn't causation. Knowing that R. gnavus bloomed during flares was like knowing that firefighters show up at fires. Was the bacterium causing the inflammation, or was it just thriving in an already inflamed environment? This chicken-and-egg problem plagued microbiome research for years. Hundreds of papers documented associations between specific bacteria and diseases, but almost none could identify a specific molecular mechanism.
The Henke discovery broke through that wall. By isolating the glucorhamnan and demonstrating its direct effect on immune cells, the researchers moved from "this bacterium is associated with Crohn's" to "this molecule from this bacterium causes this immune response." Just as the discovery of Helicobacter pylori transformed our understanding of stomach ulcers from "stress disease" to "bacterial infection," R. gnavus research is transforming how we think about inflammatory bowel disease.
The shift from "your microbiome is imbalanced" to "this specific molecule triggers this specific immune response" represents one of the most important methodological advances in IBD research in decades.
Here's where it gets biochemically weird, and genuinely surprising.
TLR4 is one of your immune system's oldest alarm bells. It sits on the surface of dendritic cells, macrophages, and monocytes, scanning for signs of bacterial invasion. Specifically, TLR4 evolved to detect lipopolysaccharide (LPS), a molecule found in the outer membrane of gram-negative bacteria. When TLR4 grabs LPS, it triggers a cascade through the MyD88-dependent pathway that produces TNF-alpha, interleukin-6, and other inflammatory molecules. This is supposed to happen when pathogens like E. coli or Salmonella breach the gut lining.
But R. gnavus is gram-positive. It shouldn't be activating TLR4 at all. The fact that its glucorhamnan polysaccharide can trigger TLR4 challenges decades of immunological assumptions about what this receptor recognizes. It's a bit like discovering that your home security system, designed to detect broken windows, also goes off when someone rings the doorbell in a specific pattern.
This finding has implications beyond Crohn's. If gram-positive bacterial sugars can activate TLR4, the known ligand repertoire for innate immune sensing is broader than anyone thought. Other gut commensals might produce similar molecules. Other autoimmune conditions might involve analogous mechanisms. The R. gnavus discovery is opening doors across immunology.
One of the most fascinating complications is that R. gnavus isn't uniformly dangerous. It lives in the guts of healthy people too, usually at low abundance without causing problems. The difference comes down to strain-level variation.
Recent research has shown that capsular polysaccharide (CPS) production varies dramatically between strains. Strains that produce CPS are actually better colonizers but paradoxically less inflammatory. CPS-producing strains are more prevalent in healthy individuals, while strains lacking CPS genes dominate in Crohn's patients and produce more TNF-alpha in laboratory experiments.
This means that standard microbiome sequencing, which identifies bacteria at the species level, misses the point entirely. You could have abundant R. gnavus in your gut and be perfectly fine, because your strains are the CPS-producing, non-inflammatory type. Functional metabolite profiling or strain-resolved genomics would be needed to distinguish friend from foe.

Adding another layer, researchers recently discovered that some R. gnavus strains produce a broad-spectrum bacteriocin called mediterrocin, an antimicrobial peptide that kills other bacteria. Disease-associated strains of R. gnavus are most susceptible to mediterrocin, raising the tantalizing possibility that the gut's own microbial warfare could be harnessed therapeutically.
"CPS production is crucial for competitive fitness during colonization in germ-free mouse intestines and is inversely correlated with the inflammatory activity of R. gnavus."
- Capsular polysaccharide study, ResearchGate
The TNF-alpha connection is particularly revealing when you look at how we currently treat Crohn's disease. Anti-TNF biologics like infliximab and adalimumab are the frontline therapies for moderate-to-severe Crohn's. A large prospective cohort study found that about 74% of Crohn's patients initially respond to anti-TNF treatment, and roughly 48% achieve clinical remission. But here's the problem: approximately 25% of patients lose their response within a few years.
These drugs work downstream, mopping up TNF-alpha after it's already been produced. They don't address why the TNF-alpha is being produced in the first place. If R. gnavus glucorhamnan is continuously activating TLR4 and triggering fresh waves of TNF-alpha, you're essentially bailing water while the hole in the boat keeps growing. An upstream approach that targets the bacterial polysaccharide itself could be far more effective and less immunosuppressive.
Current anti-TNF drugs mop up inflammation after the fact. Targeting the bacterial sugar that triggers it could be like fixing the leak instead of just bailing water.
The race to translate microbiome science into therapy is genuinely global, and different research cultures are approaching the problem from different angles.
In the United States, the focus has been on molecular mechanism and drug target identification, exemplified by the Clardy lab's work at Harvard and NIH-funded research programs. American researchers tend to think in terms of small-molecule drugs or biologics that could block the glucorhamnan-TLR4 interaction.
European researchers, particularly at the Quadram Institute in the UK where Nathalie Juge's lab operates, have focused on understanding R. gnavus's ecological niche. Juge's team discovered that R. gnavus uses a specialized enzyme called intramolecular trans-sialidase to harvest sialic acid from gut mucus, and that it possesses a unique transporter for the rare sugar 2,7-anhydro-Neu5Ac that no other gut microbe can use. This ecological approach suggests dietary or prebiotic strategies could starve inflammatory strains of their preferred nutrients.

Meanwhile, research teams across Asia and Europe are investigating fecal microbiota transplantation (FMT) as a therapeutic strategy. Germ-free mouse experiments have shown that transplanting microbiota from Crohn's patients results in enrichment of R. gnavus and development of colitis, providing direct evidence that the bacterium can initiate disease. These models are being used to test whether curated microbial communities lacking inflammatory R. gnavus strains could prevent or reduce flares.
The development of genetic toolkits for manipulating R. gnavus, including shuttle vectors and gene deletion systems, represents another frontier. After nearly a decade of failed attempts, researchers can now knock out specific genes and observe the effects, enabling precise dissection of which molecular components drive colonization versus inflammation.
Within the next decade, the practical implications of this research could reach your doctor's office in several ways. Stool-based diagnostic tests that measure R. gnavus glucorhamnan levels could provide early warning of impending flares, giving patients and clinicians time to intervene preventively. Precision antibiotics or bacteriocin-based therapies that selectively target inflammatory strains while leaving beneficial bacteria intact could replace the blunt-instrument immunosuppressants currently in use.
For Crohn's patients and their caregivers, the most important takeaway is this: the disease isn't random. There's a specific molecular pathway connecting a specific bacterial product to the immune overreaction that causes flares. That specificity is exactly what makes it targetable. The era of "your gut bacteria are out of balance" hand-waving is giving way to something far more precise, far more testable, and far more hopeful. The sugar that triggers the storm has been found. Now the work of stopping it can truly begin.

Saturn's moon Titan may harbour liquid water beneath its frozen crust, kept from freezing by ammonia acting as a natural antifreeze. New Cassini data suggests the interior could be slush with warm water pockets rather than a global ocean, and NASA's Dragonfly mission launching in 2028 aims to investigate whether this exotic environment could support life.

Bacteroides thetaiotaomicron uses 88 specialized gene clusters and over 260 enzymes to decode and digest dietary fibers humans can't break down, converting them into essential short-chain fatty acids. When fiber runs out, it eats your gut's protective mucus instead, with cascading health consequences.

Scientists are restoring Ice Age ecological dynamics through rewilding projects like Siberia's Pleistocene Park and de-extinction efforts by Colossal Biosciences. These initiatives aim to reintroduce megafauna or their proxies to repair broken ecosystems, protect Arctic permafrost, and slow climate change.

The cheerleader effect is a proven cognitive bias where people look more attractive in groups because the brain automatically averages faces, smoothing out individual flaws. Research shows the sweet spot is 3-5 people, it works for all genders, and it has real implications for dating apps and social media strategy.
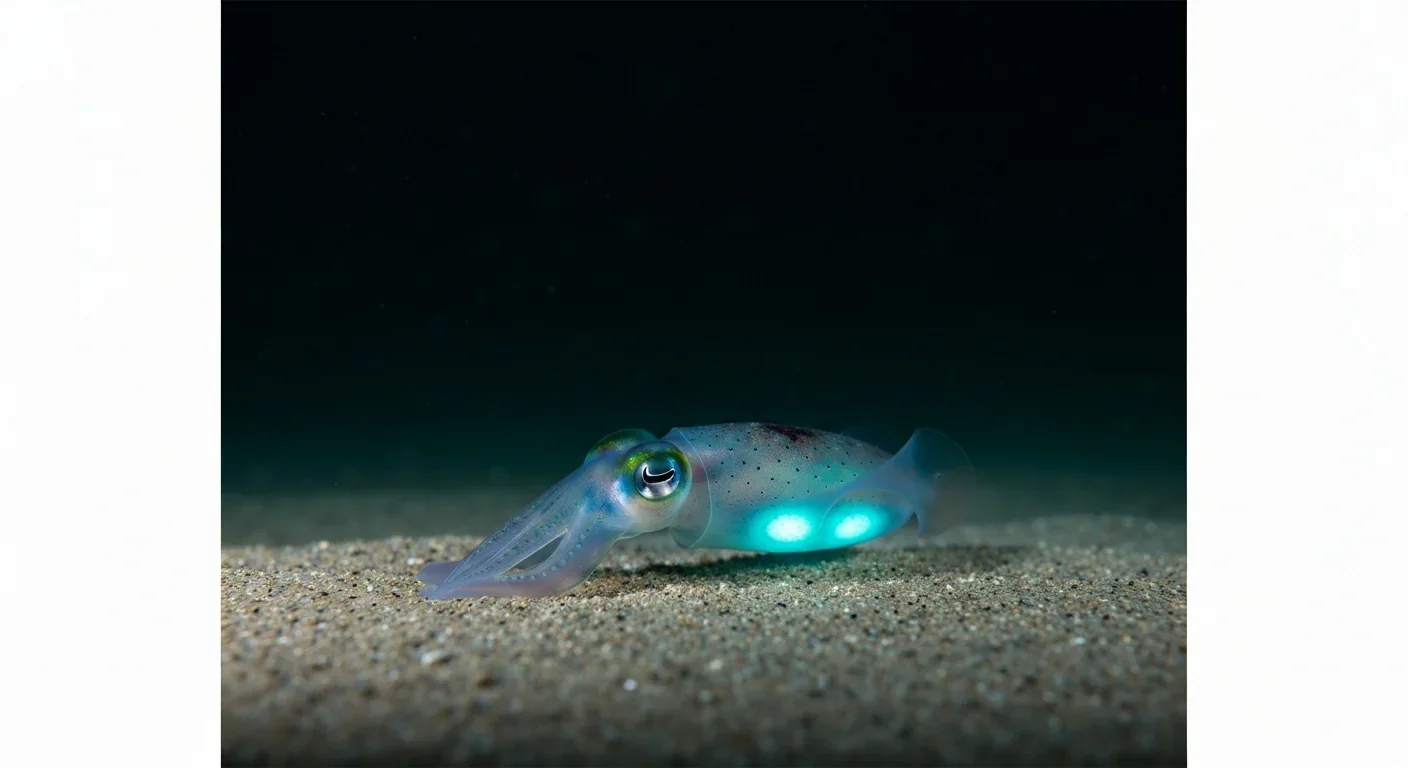

The Hawaiian bobtail squid farms bioluminescent bacteria in a specialized light organ to erase its shadow in moonlit waters. This partnership, where bacteria reshape the squid's body and communicate through quorum sensing, is teaching scientists how host-microbe relationships work and inspiring new medical and biotech applications.

Millions are leaving social media platforms driven by privacy scandals, mental health concerns, and algorithmic manipulation. While 60% relapse within a week, those who stay away report dramatically improved wellbeing, and decentralized alternatives like Bluesky are surging.

Building a reliable quantum computer requires roughly 1,000 fragile physical qubits per logical qubit due to surface code error correction overhead. New code families like LDPC and neutral-atom platforms are racing to slash that ratio, with some teams claiming it could drop to as few as 5-to-1.